import stistools
from astropy.io import fits
Defringing STIS CCD Spectra With The Stistools Defringing Tool Suite¶
STIS CCD spectra taken at wavelengths >7000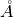 (the
G750M and G750L modes) are impacted by “fringing”, a phenomenon caused
by interference of multiple reflections between the two surfaces of the
CCD in cases where the wavelength of the incident light is a small
integer multiple of the distance between the two surfaces of the CCD
(Goudfrooij et al. 1998).
(the
G750M and G750L modes) are impacted by “fringing”, a phenomenon caused
by interference of multiple reflections between the two surfaces of the
CCD in cases where the wavelength of the incident light is a small
integer multiple of the distance between the two surfaces of the CCD
(Goudfrooij et al. 1998). stistools contains four tools used for
correcting the fringing effect, normspflat, prepspec,
mkfringeflat, and defringe, which are located in the
stistools.defringe package. These tools are python ports of the
original IRAF/PyRAF stis defringing tools. The following serves as a
guide for using these tools to defringe STIS CCD data, and presents
several practical examples for common use cases.
This guide assumes that the stistools package has been installed.
For instructions on how to install stistools, consult the Getting
Started Guide:
https://stistools.readthedocs.io/en/latest/gettingstarted.html.
Additionally, it assumes you’ve setup CRDS which is also walked through
in the Getting Started Guide.
Data Setup¶
To defringe a STIS CCD G750M/G750L observation, the fits files for the
fringed science image in addition to the associated contemporaneous
fringe flat for the observation are required. It’s standard practice to
take contemporaneous fringe flats for G750M/G750L spectra, the filename
for the fringe flat is stored in the primary header of the science
image, specifically in the 'FRNGFLAT' keyword. Additionally, the
'OPT_ELEM' keyword in the primary header contains the mode used for
the observation, which will be important as there are a few differences
in the defringing process depending on the mode. These differences will
be highlighted in each step.
#setup data paths
sci_file = "odvkl3050" # the science file
flat_file = "odvkl3080" # the associated fringe flat
#Confirm that the flat file is indeed the associated fringe flat for odvkl3050
print("Associated Fringe Flat: "+fits.getheader(f"{sci_file}_raw.fits",0)['FRNGFLAT'])
print("Observing Mode: "+fits.getheader(f"{sci_file}_raw.fits",0)['OPT_ELEM'])
Associated Fringe Flat: ODVKL3080
Observing Mode: G750L
1. Normalize the Fringe Flat¶
The normspflat tool is used to normalize the contemporaneous flat
field image. The do_cal parameter tells normspflat whether to
calibrate the flat file using calstis or not. If using a raw input
image, do_cal should be set to true, if using a calibrated image
(crj or sx2) do_cal can be set to false. The output file,
which has the nsp identifier in our example, is the normalized flat
field for use in the next steps. It’s worth noting that in the
do_cal=True case, normspflat will print out a message
identifying the most calibrated output file produced by calstis, which
in this case is a crj file.
stistools.defringe.normspflat(f"{flat_file}_raw.fits",
f"{flat_file}_nsp.fits", do_cal=True,
wavecal=f"{sci_file}_wav.fits")
File written: /Users/stisuser/data/path/odvkl3080_crj.fits
G750M/G750L Point of Difference: Fringe flat images taken with G750L include not only the IR fringing at wavelengths greater than 7500 Angstroms, but also some fringes at wavelengths less than 6000 Angstroms due to an order-sorter filter. Since these order-sorter fringes are already included in the sensitivity function, they should not be included in the fringe flat, and so these columns should be set to unity in the normalized fringe flat. The following code accomplishes this:
# Flatten the blue end of the flat-field image [ONLY FOR G750L]
with fits.open(f"{flat_file}_nsp.fits", mode='update') as hdulist:
hdulist[1].data[:,:250] = 1
2. Prepare the Science File for the Defringing Correction (Optional)¶
The prepspec tool is used to calibrate the raw science image. It
essentially runs the science through calstis with a specific set of
calibration flags. prepspec will not overwrite image products, so be
sure to remove any higher level science products (flt, crj,
sx1, sx2) from the working directory before you run it.
Note: Running prepspec is an optional step, if you already have
calibrated science data (crj/sx2) then running prepspec is
not essential. The main purpose of prepspec is to run calstis with a
specific set of calibration flags turned on (e.g. the keywords in the
header that control which calibration steps are performed and omitted by
calstis during calibration). In the typical case, the default calstis
flags will be sufficient for defringing. However, you may wish to delete
these data products and rebuild from the raw science file with
prepspec to ensure that the correct calibration was done on the
files if you are uncertain.
%%capture
#capture the long calstis output
stistools.defringe.prepspec(f"{sci_file}_raw.fits")
3. Match Fringes in the Fringe Flat Field and the Science Spectra¶
The mkfringeflat tool is used to calculate the appropriate shifts
and scale factors needed to match the fringes in the fringe flat and the
science spectra. The output is a shifted and scaled fringe flat which
can be named however you wish, but we refer to as an frr file
product in our documentation. The best shift and scale factors are
obtained by finding the values that minimize the RMS within a
user-specified search range. The parameters that control the range and
step size for the shift and scale have default values (shown explicitly
below) that should serve most use cases well. mkfringeflat will warn
the user if the best shift and scale values were found at the edge of
the range, suggesting the range may need to be expanded further to find
the best values. The beg_shift and end_shift arguments can be
used to adjust the shift range, while the beg_scale and
end_scale arguments can be used to adjust the scale range.
G750M/G750L Point of Difference: The appropriate file type to use as
the input science file depends on the observation mode. For G750L,
crj files should be used. For G750M, geometric correction is
required before defringing can take place, so sx2 products should be
used.
# choose the correct science product type based on the mode
mode = fits.getheader(f"{sci_file}_raw.fits",0)['OPT_ELEM']
if mode == "G750L":
prod_type = "crj"
elif mode == "G750M":
prod_type = "sx2"
stistools.defringe.mkfringeflat(f"{sci_file}_{prod_type}.fits", f"{flat_file}_nsp.fits",
f"{flat_file}_frr.fits", beg_shift=-0.5, end_shift=2, shift_step=0.1,
beg_scale=0.8, end_scale=1.7, scale_step=0.04)
mkfringeflat.py version 0.1
- matching fringes in a flatfield to those in science data
Extraction center: row 583
Extraction size: 11.0 pixels [Aperture: 52X2]
Range to be normalized: [578:589,4:1020]
Determining best shift for fringe flat
shift = -0.500, rms = 8.8683
shift = -0.400, rms = 9.5374
shift = -0.300, rms = 10.3134
shift = -0.200, rms = 10.8593
shift = -0.100, rms = 12.8151
shift = 0.000, rms = 2.8657
shift = 0.100, rms = 2.9300
shift = 0.200, rms = 2.9326
shift = 0.300, rms = 3.0001
shift = 0.400, rms = 3.0489
shift = 0.500, rms = 3.0998
shift = 0.600, rms = 3.1530
shift = 0.700, rms = 3.2087
shift = 0.800, rms = 3.2670
shift = 0.900, rms = 3.3279
shift = 1.000, rms = 3.3917
shift = 1.100, rms = 3.9375
shift = 1.200, rms = 8.4936
shift = 1.300, rms = 2.5887
shift = 1.400, rms = 2.7323
shift = 1.500, rms = 2.9274
shift = 1.600, rms = 3.2250
shift = 1.700, rms = 3.7717
shift = 1.800, rms = 5.1464
shift = 1.900, rms = 12.0936
shift = 2.000, rms = 3.4937
Best shift : 1.347 pixels
Shifted flat : odvkl3080_nsp_sh.fits
(Can be used as input flat for next iteration)
Determining best scaling of amplitude of fringes in flat
Fringes scaled 0.800: RMS = 2.7298
Fringes scaled 0.840: RMS = 2.7122
Fringes scaled 0.880: RMS = 2.6956
Fringes scaled 0.920: RMS = 2.6800
Fringes scaled 0.960: RMS = 2.6653
Fringes scaled 1.000: RMS = 2.6515
Fringes scaled 1.040: RMS = 2.6384
Fringes scaled 1.080: RMS = 2.6260
Fringes scaled 1.120: RMS = 2.6142
Fringes scaled 1.160: RMS = 2.6031
Fringes scaled 1.200: RMS = 2.5925
Fringes scaled 1.240: RMS = 2.5825
Fringes scaled 1.280: RMS = 2.5730
Fringes scaled 1.320: RMS = 2.5639
Fringes scaled 1.360: RMS = 12.0382
Fringes scaled 1.400: RMS = 10.9230
Fringes scaled 1.440: RMS = 5.0855
Fringes scaled 1.480: RMS = 4.9331
Fringes scaled 1.520: RMS = 4.7929
Fringes scaled 1.560: RMS = 4.6632
Fringes scaled 1.600: RMS = 4.5430
Fringes scaled 1.640: RMS = 4.4316
Fringes scaled 1.680: RMS = 4.3276
Best scale : 1.284
Output flat : odvkl3080_frr.fits
(to be used as input to task 'defringe.py')
4. Defringe the Science Spectra¶
The final step is to use the defringe tool to divide the scaled and
shifted fringe flat off of the calibrated science spectra, removing the
fringing pattern.
G750M/G750L Point of Difference: As in the previous step, the input
science product type is dependent on mode. (G750L: crj, G750M:
sx2)
stistools.defringe.defringe(f"{sci_file}_{prod_type}.fits", f"{flat_file}_frr.fits", overwrite=True)
Fringe flat data were read from the primary HDU
108 pixels in the fringe flat were less than or equal to 0
Imset 1 done
Defringed science saved to odvkl3050_drj.fits
'odvkl3050_drj.fits'
We now have a drj file that is the fully defringed calibrated
science spectra. This file functionally behaves as the crj file used
to produce it, and may be worked with in the same manner.
G750M/G750L Point of Difference: If working with a G750M
observation, the output product by default will have the s2d
identifier.
Extraction of 1D Spectra from Defringed Science Products¶
As mentioned above, the defringed science products may be worked with as
normal calstis products. Typically, the next step would be extract 1D
spectra from these files. This can be accomplished by continuing the
calibration through calstis.calstis or performing the extraction
step individually using x1d.x1d, please refer to the documentation
for those tools if you’re looking for guidance on that step.
1D Extraction of G750M Spectra¶
It’s important to note that, at this time, x1d and calstis are
not able to extract 1D spectra from the G750M sx2 products. sx2
products have been geometrically rectified, which generates correlated
errors between wavelength bins. These errors are not well-handled by the
standard pipeline extraction algorithms.